Transposable Elements, Epigenetics and Evolution
Transposable elements are repetitive DNA sequences that account for close to 50% of the DNA found in the human genome. Epigenetic silencing of transposable elements ensure their tight regulation throughout development. While we still know very little about the function of transposable elements, emerging evidence shows that transposable elements are integral to the evolution of the human genome.
Have you ever wondered why corn kernels come in so many different colors? What causes the random yellow and white pattern in a cob of peaches and cream?
In the 1940s, American scientist Barbara McClintock won a Nobel Prize for figuring out the answer to similar questions. Turns out, color variation in corn kernels is caused by an exchange of genetic material, often due to transposable elements or “jumping DNA”.
Since Dr. McClintock’s experiments on corn, scientists have made great strides to learn about transposable elements—repetitive sequences of DNA make up almost half the human genome. They are common in other species as well, with different categories of transposable elements making up a different proportion of every genome. More recent research is looking at why this is the case and how transposable elements are responsible for more than the color of corn kernels. In fact, they play a vital role in species diversity, evolution and embryo development.
Jumping DNA
Transposable elements come in many lengths and forms. But they have the ability to do something other DNA sequences can’t do—they can jump from one spot in the genome to another.
Some transposable elements—the transposons—move by directly cutting themselves out of one spot and pasting themselves into another, not unlike the cut-and-paste function in a word processing system. Other transposable elements—the retrotransposons—move through something more like a copy-and-paste function, where the original DNA is copied into RNA and then pasted into a new spot in the genome through transcription.
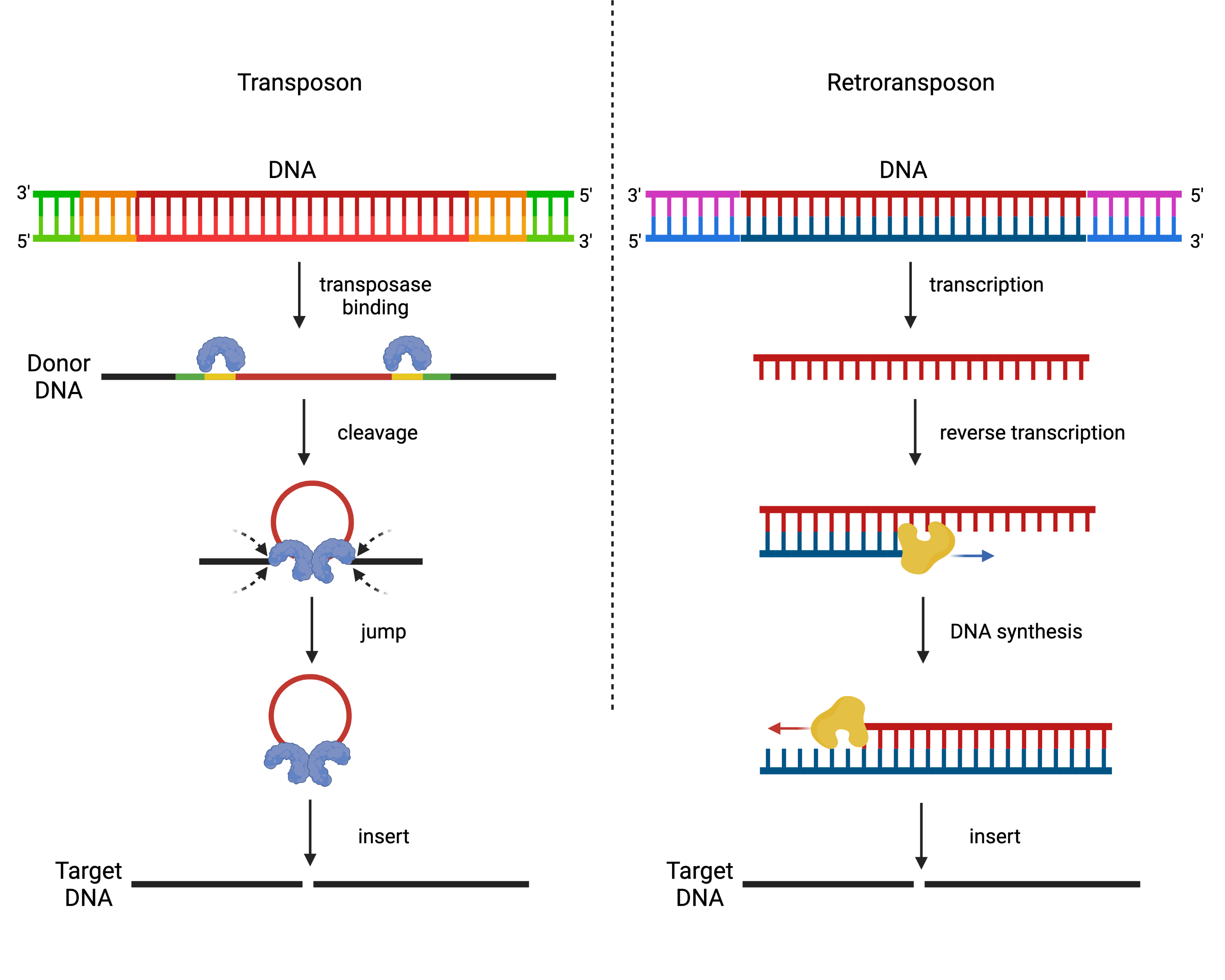
(Image created in BioRender.com)
Effect on the genome
For a long time, scientists didn’t really understand the purpose of transposable elements. They were considered “selfish” and viewed as somewhat parasitic. The thinking was that all this jumping around was likely to disrupt genes with negative consequences. They have, for example, been associated with complex human disease such as cancer and schizophrenia.
When the epigenetic silencing of transposable elements was discovered, it supported this theory. Transposable elements needed to be silenced—or in a sense grounded, so they could no longer jump—in order to prevent genomic chaos that could lead to disease.
Epigenetic silencing
Epigenetic modifications to the genome are biochemical changes to the DNA, usually thought to regulate gene expression. Silencing of a gene is the equivalent of “turning it off”. To flip the switch, a chemical group can be added to the DNA such as a methyl group resulting in DNA methylation, which prevents it from being read by the cell. Similarly, DNA that’s tightly wrapped around histones can’t be read either.
In order to jump, transposable elements need to be read. The silencing of transposons limits the transcription of transposase—the enzyme needed to cut and paste a transposon from one part of the genome to another. For retrotransposons, silencing prevents them from performing the copy function. In addition to epigenetic modification, regulatory control over transposable elements is accomplished in other ways as well.
Jumping with purpose
While it’s true that transposable elements can be disruptive—and therefore need to be “silenced”—there are three problems with keeping jumping DNA down.
First, transposable elements tend to be located in areas of the genome that have a lot of genes. And when the transposable elements are silenced, the epigenetic modification tends to spread to those neighboring genes—which may or may not cause problems for the host.
Second, it’s been shown that transposable elements are responsible for creating new genes associated with species variability and evolution. They essentially accomplish this by fusing different parts of a gene together and inserting their sequence in between. This is known as exon shuffling and it’s a bit like rearranging Lego blocks to create different objects.
Third, scientists have shown that certain transposable elements are required to turn genes on and off during embryo development. This results in variation between gene expression in different cells and different times, allowing for different tissues to form from the same DNA or genome.
It’s all about balance
There’s still more to learn about transposable elements and the role they play in evolution and human development. But research is re-framing our thinking such that transposable elements can no longer be seen as selfish, parasitic disruptors.
Intriguingly, it may be that transposable elements—and the epigenetic silencing of those transposable elements—have created the genomic change necessary for evolution to occur. They play a key role in gene expression, which influences developmental differentiation and physiology. In other words, they help make different species… different.
Learn More
- Learn more about the role of epigenetic silencing of transposable elements in evolution here.
- Learn more about current epigenetic studies related to transposons here.
- Find out ten things about transposable elements here.
- Read more about the different classes of transposable elements here.
We would like to thank Dr. Mathieu Lupien from Princess Margaret Cancer Centre for reviewing this piece.

